|
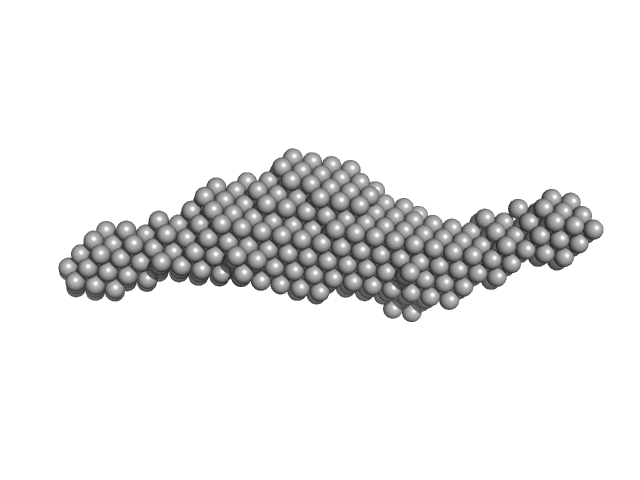
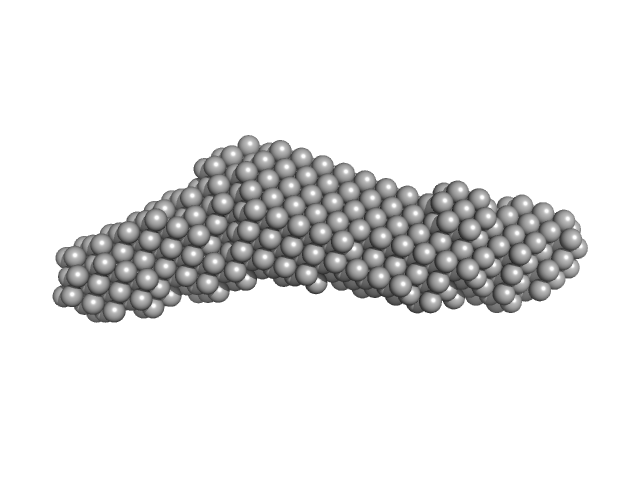
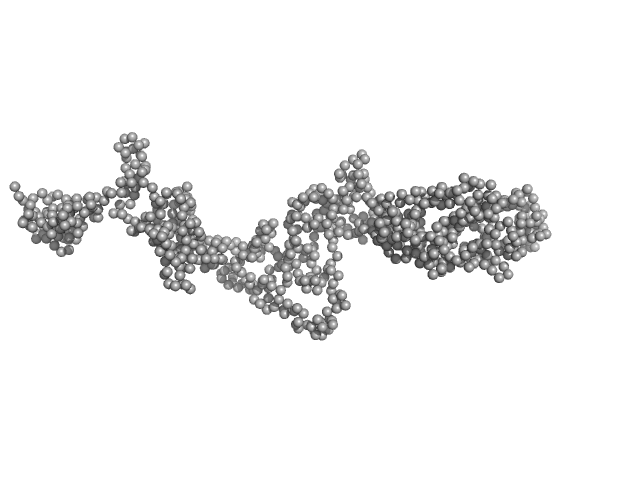
Synchrotron SAXS data from solutions of the autoinhibited dimer of Cytohesin-3 (Grp1, amino acids 14-399) in 20 mM Tris, 150 mM NaCl, 2 mM MgCl2, 0.1% 2-mercaptoethanol, 5% glycerol, 0.001 mM insitol 1,3,4,5-tetrakis phosphate, pH 8 were collected on the BioCAT 18ID beam line at the Advanced Photon Source (APS), Argonne National Laboratory (Lemont, IL, USA) using a MAR 165 CCD detector at a sample-detector distance of 2.5 m and at a wavelength of λ = 0.10332 nm (I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). In-line size-exclusion chromatography (SEC) SAS was employed. The SEC parameters were as follows: A 100.00 μl sample at 12 mg/ml was injected at a 0.25 ml/min flow rate onto a GE Superdex 200 Increase 3.2/300 column at 20°C. SAXS data sets were acquired with 1 s exposures at 5 s intervals during the elution. Raw SAXS images were radially averaged on a log scale over the s range 0.0621-3.33 nm-1 and normalized by the incident beam intensity. The protein scattering profile was reconstructed by singular value decomposition and linear combination (SVD-LC) as described in Malaby et al. (2015) Methods for analysis of size-exclusion chromatography-small angle X-ray scattering and reconstruction of protein scattering. J Appl Crystallogr 48: 1102-1113.
The protein was equilibrated with a 1.2 molar excess of inositol 1,3,4,5-tetrakis phosphate (IP4) and concentrated to 12 mg/ml prior to injection.
|
|
 s, nm-1
s, nm-1