|
Synchrotron SAXS data from solutions of aromatic-L-amino-acid decarboxylase (L353P) bound to pyridoxal 5'-phosphate in 50 mM HEPES, pH 7.4 were collected on the B21 beam line at the Diamond Light Source (Didcot, UK) using a Eiger 4M detector at a sample-detector distance of 3.7 m and at a wavelength of λ = 0.1 nm (I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). In-line size-exclusion chromatography (SEC) SAS was employed. The SEC parameters were as follows: A 50.00 μl sample at 4 mg/ml was injected at a 0.16 ml/min flow rate onto a Shodex KW403-4F column at 20°C. 620 successive 3 second frames were collected. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted.
Pyridoxal 5'-phosphate: https://www.ebi.ac.uk/chebi/searchId.do?chebiId=CHEBI:18405. CAUTION: Data used to calculate p(r) have been significantly truncated at both low- and high-s.
|
|
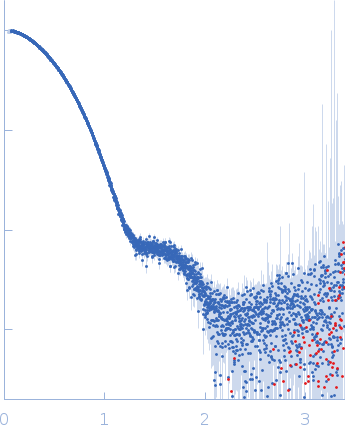 s, nm-1
s, nm-1
