A shape-shifting redox foldase contributes to Proteus mirabilis copper resistance.
Furlong EJ,
Lo AW,
Kurth F,
Premkumar L,
Totsika M,
Achard MES,
Halili MA,
Heras B,
Whitten AE,
Choudhury HG,
Schembri MA,
Martin JL
Nat Commun
8:16065
(2017 Jul 19)
|
|
|
|
|
| Sample: |
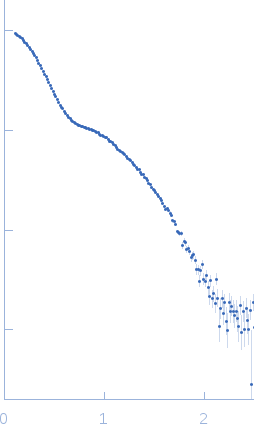
C-terminal catalytic domain of Suppressor of Copper Sensitivity C protein monomer, 20 kDa Proteus mirabilis protein
|
| Buffer: |
25 mM HEPES 150mM NaCl 1mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2012 Feb 29
|
|
| RgGuinier |
3.7 |
nm |
| Dmax |
10.5 |
nm |
| VolumePorod |
92 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
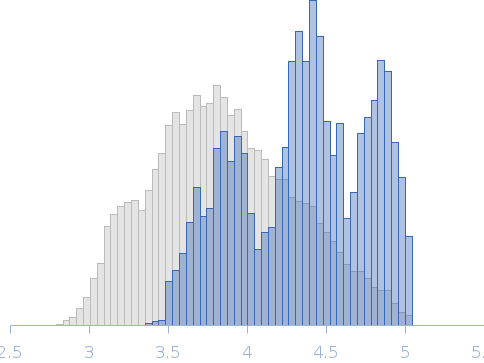
Suppressor of Copper Sensitivity C protein (mutant) trimer, 73 kDa Proteus mirabilis protein
|
| Buffer: |
25 mM HEPES 150mM NaCl, 1mM DTT,, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2016 Jul 22
|
|
| RgGuinier |
4.4 |
nm |
| Dmax |
13.5 |
nm |
| VolumePorod |
108 |
nm3 |
|
|