|
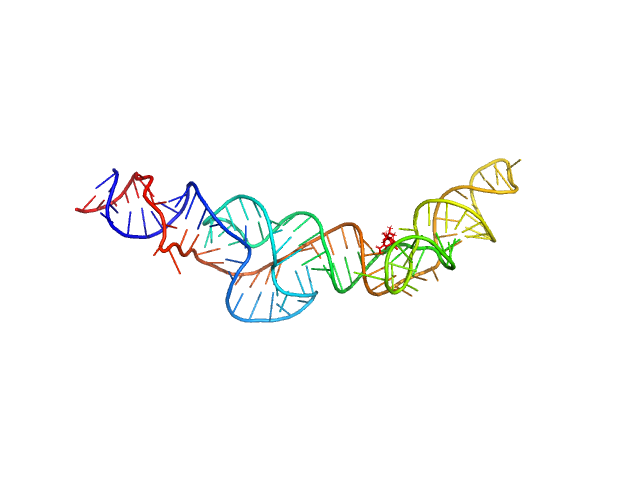
Synchrotron SAXS data from solutions of holo-D43 in 10 mM Bis-Tris, 100 mM KCl, and 1 mM MgCl2, pH 6.8 were collected on the 16-ID (LiX) beam line at the National Synchrotron Light Source II (NSLS-II; Upton, NY, USA) using a Pilatus3 S 1M detector at a sample-detector distance of 3.7 m and at a wavelength of λ = 0.08202 nm (I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). Solute concentrations ranging between 1.2 and 10 mg/ml were measured at 20°C. 20 successive 0.500 second frames were collected. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted. The low angle data collected at lower concentrations were extrapolated to infinite dilution and merged with the higher concentration data to yield the final composite scattering curve.
The MW for the RNA was calculate using the volume of correlation method.
|
|
 s, nm-1
s, nm-1