|
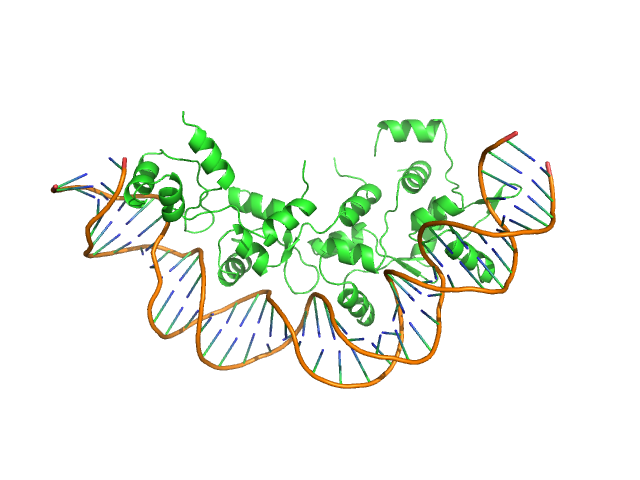
Synchrotron SAXS data from solutions of His-tagged RdfS excisionase bound to dsDNA 40mer attP_8 in 150 mM Tris-HCl, 300 mM NaCl, 5% v/v glycerol, pH 7.4 were collected on the SAXS/WAXS beam line at the Australian Synchrotron (Melbourne, Australia) using a Pilatus 1M detector at a sample-detector distance of 2.6 m and at a wavelength of λ = 0.103 nm (I(s) vs s, where s = 4πsinθ/λ, and 2θ is the scattering angle). In-line size-exclusion chromatography (SEC) SAS was employed. The SEC parameters were as follows: A 50.00 μl samples at 6H-RdfS at 390 µM, dsDNA attP_8 at 500 µM was injected at a 0.30 ml/min flow rate onto a GE Superdex 200 Increase 5/150 column at 23°C. 36 successive 1 second data frames were selected from the SEC sample peak (from a total of 800 individual frames measured from the entire SEC elution) and processed to generate the profile displayed in this entry. The data were normalized to the intensity of the transmitted beam and radially averaged; the scattering of the solvent-blank was subtracted.
|
|
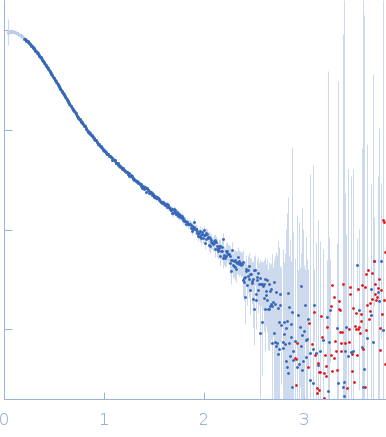 s, nm-1
s, nm-1